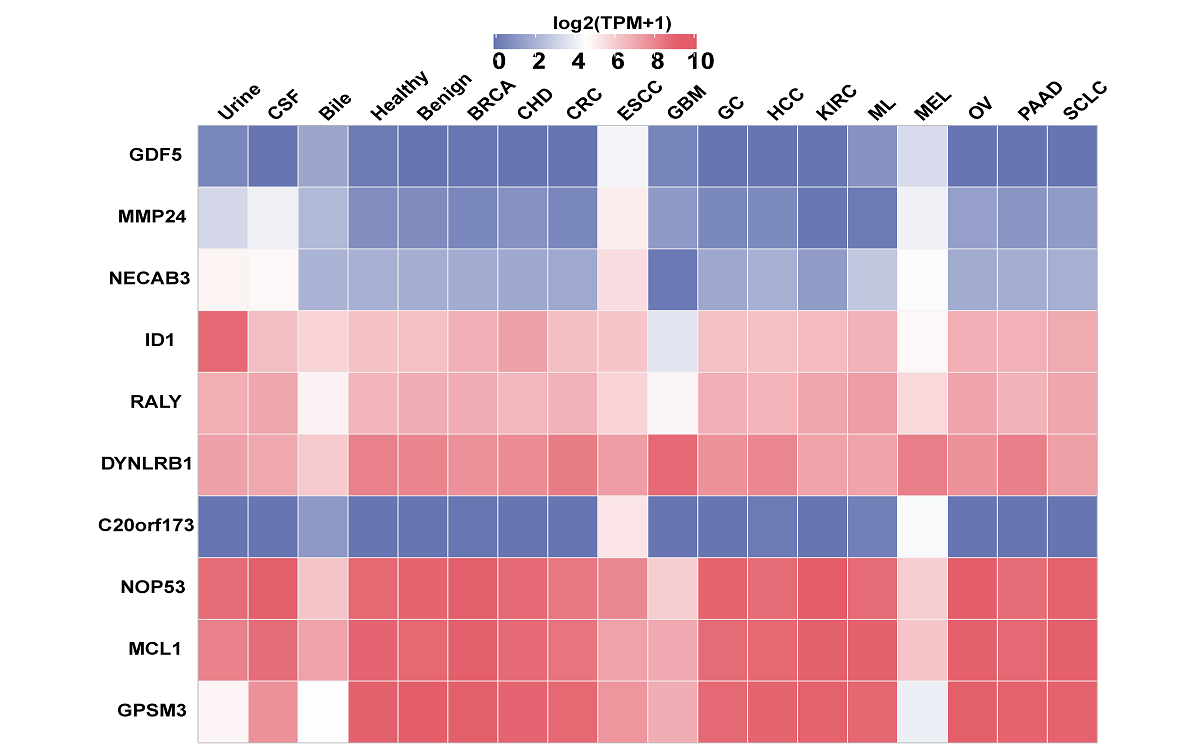
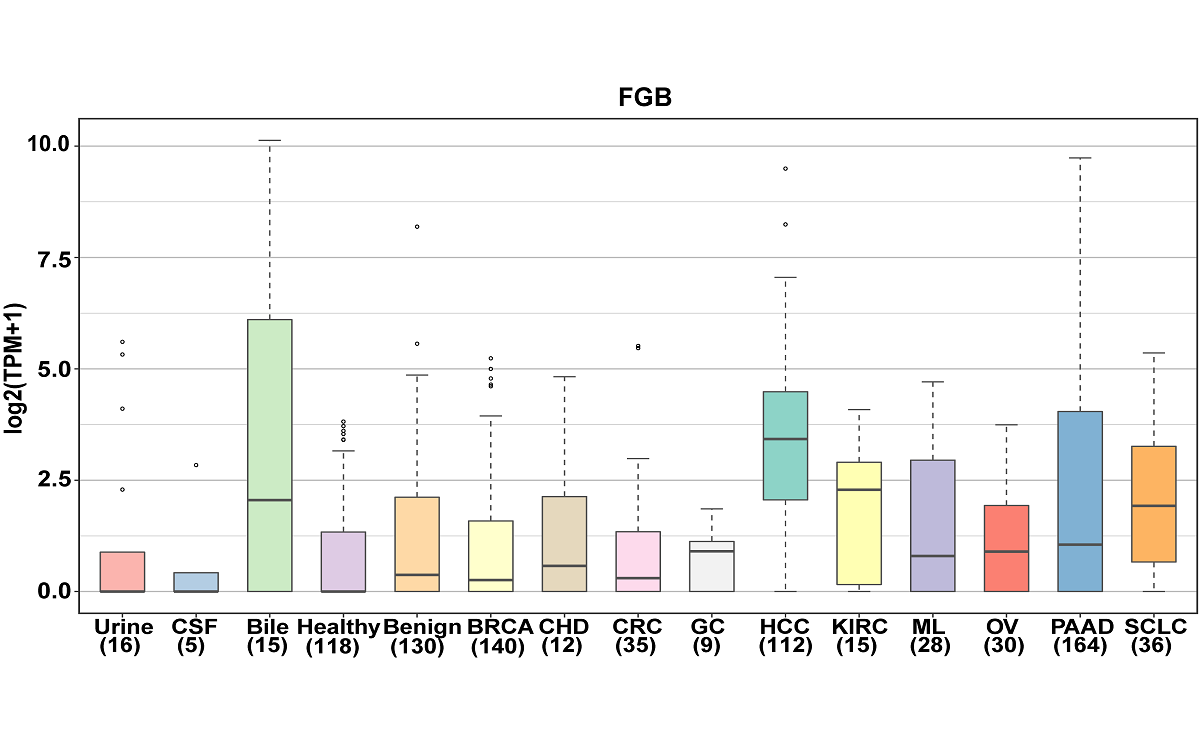
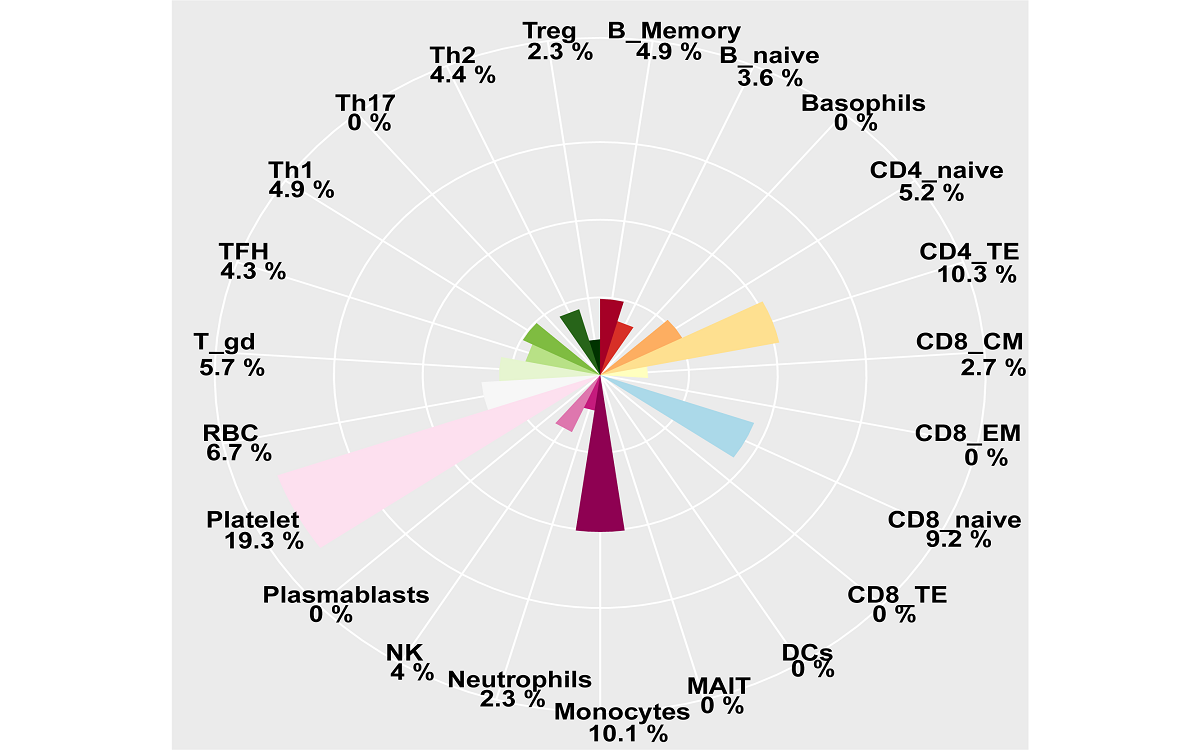
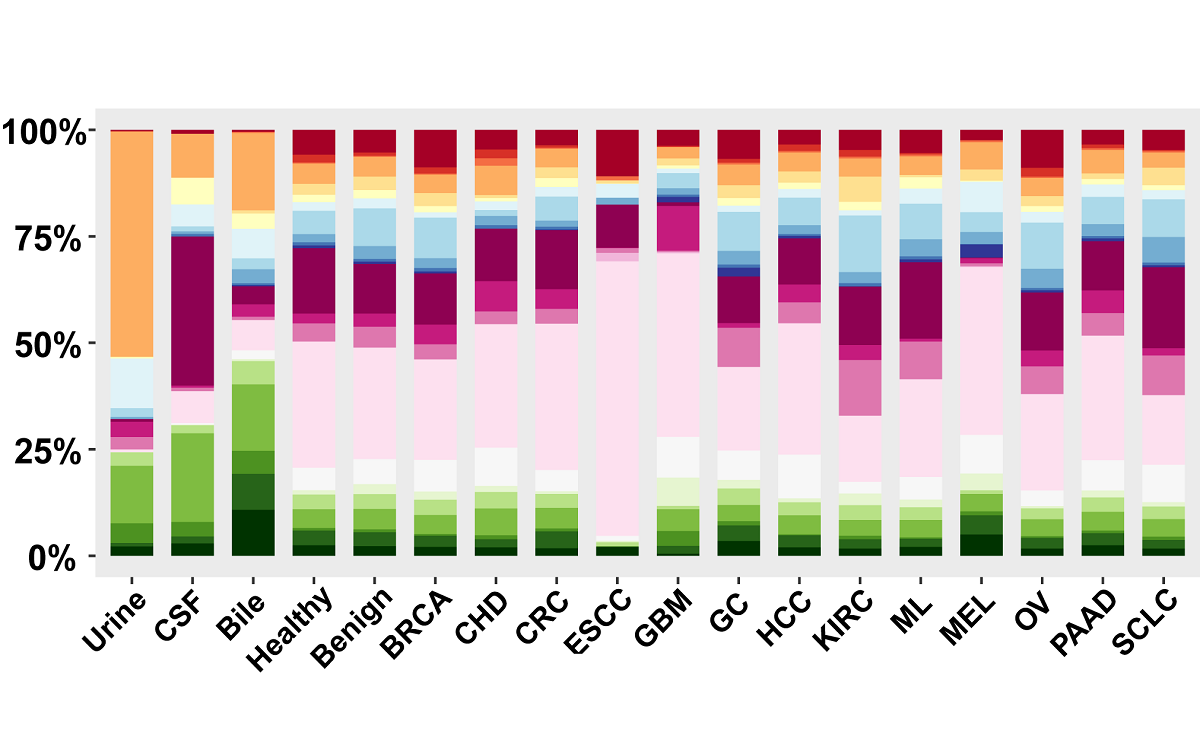
mRNA, lncRNA and circRNA in extracellular vesicles
Welcome to exoRBase 2.0
exoRBase is a repository of extracellular vesicles (EVs) long RNAs (exLRs) derived from RNA-seq data analyses in different human body fluids. The exLRs contain messenger RNA (mRNA), long non-coding RNA (lncRNA), and circular RNA (circRNA).
Here, we have updated the exoRBase database to version 2.0 by collecting about 1,000 RNA-seq data of EVs from four types of human body fluids. exoRBase 2.0 features the integration and visualization of RNA expression profiles, as well as the functional pathway-level changes and the heterogeneity of circulating-EVs origins.
exoRBase 2.0 provides the comprehensive annotation and expression landscapes of exLRs. It will facilitate the identification of novel exLR signatures from human body fluids and will help discover new circulating biomarkers for the improvement of tumor diagnosis and therapy.
Release Notes&News
- exoRBase 2.0 is published in Nucleic Acids Research. [November 2021] [ Full Text]
- exoRBase has been updated to verson 2.0. [August, 2021] [ Version 1.0]
- exoRBase has received more than 70,000 universal visitors. [June, 2021]
- EV-origin: enumerating the tissue-cellular origin of circulating extracellular vesicles using exLR profile, published in Comput Struct Biotechnol J. [October 2020] [ Full Text]
- The exLR biomarkers used for the detection of pancreatic ductal adenocarcinoma published in Gut. [March 2020] [ Full Text]
- The feature and application of human blood exLRs published in Clin Chem. [March 2019] [ Full Text]
- exoRBase has received 10,000 visitors. [May, 16 2018]
- exoRBase is published in Database Issue of Nucleic Acids Research. [October 2017] [ Full Text]
- exoRBase is publicly accessible. [August 2017]
- 77 experimental validations from 18 published literatures have been included. [July 2017]
- GTEx and circBase data have been used to annotate possible original cells/tissues of exosomal RNAs. [July 2017]
- 92 RNA-seq samples from seven experiments have been integrated into exoRBase. [June 2017]